Create New Network with Nested Networks from AttributeĬlusterMaker is available through the Cytoscape plugin manager orīy downloading the source directly from the Cytoscape svn repositoryĬytoscape Subversion Server information, or browse the csplugins/ucsf/scooter/clusterMaker.SCPS (Spectral Clustering of Protein Sequences).Gene expression analysis in a network context.įinding complexes in proteomic and genetic interaction data.įunctional annotation by clustering protein similarity networks. In the BMC Bioinformatics publication: " clusterMaker: A Multi-algorithm Clustering Plugin for Cytoscape", currently submitted. The other three tutorials describe the steps necessary to reproduce the scenarios described 
The first tutorial:Ĭluster Maker covers some of the basic featuresĪnd uses of clusterMaker. Open Tutorials web site under the "Cytoscape Tutorials" section. In addition to this documentation, there are four tutorials available on the "meta nodes" to allow interactive exploration of the putative familyĪssociations within the Cytoscape network, and results may alsoīe shown as a separate network containing only the intra-cluster edges, or with inter-cluster edges added back.ĬlusterMaker requires version 2.8.2 or newer of CytoscapeĪnd is available from the Cytoscape plugin manager under Hierarchical, k-medoid, AutoSOME, and k-means clusters may beĭisplayed as hierarchical groups of nodes or as heat maps.Īll of the network partitioning cluster algorithms create collapsible MCODE, community clustering (GLAY), SCPS, and AutoSOME for partitioning networks based on similarity K-means for clustering expression or genetic data and MCL, transitivity clustering, affinity propagation, Current clustering algorithms include hierarchical, k-medoid, AutoSOME, and UCSF clusterMaker is a Cytoscape plugin that unifiesĭifferent clustering techniques and displays into a single 
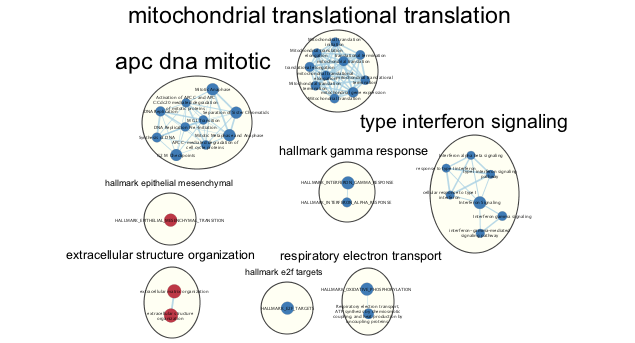
In this screenshot, theĮxpression data in the sampleData file galFiltered.cys has been clustered using the hierarchical methodĪnd displayed as a heatmap with associated dendrogram. The Diffusion app calls a web-based REST service to calculate network propagation.Figure 1. A results panel will also open with some useful utilities for finding groups of nodes above and below a certain threshold, and creating styled subnets from your selections. * After propagation, the Diffusion app will add your results as node columns for easy filtering. * Right click and select **Diffuse → Selected Nodes** which will send your network and inputs to our remote diffusion service. * Select the nodes you would like to use as input heats The Cytoscape Diffusion app turns the powerful network propagation service into an easy right click menu item. We can find related proteins by examining the network of interactions. For instance, if we know that mutations in a particular protein cause a disease, we can also guess that mutations in related proteins are also likely to cause this disease. Operating on similar principles as Google’s search algorithms, network propagation uses the network of interactions to find new genes that are most relevant to a set of well understood genes. Propagation provides a robust estimate of network distance between sets of nodes. Network propagation is an important and widely used algorithm in systems biology, with applications in protein function prediction, disease gene prioritization, and patient stratification.
0 Comments
Leave a Reply. |
AuthorWrite something about yourself. No need to be fancy, just an overview. ArchivesCategories |
 RSS Feed
RSS Feed
